by Natasha Berthold
Natasha Berthold is a 2022 CEED Summer Research Fellow, although for her it’s a Winter Research Fellowship as she is a PhD student at the University of Western Australia, in Perth, Australia.
In 2019, the eating disorders research field took a significant step forward: a global study identified eight regions across the human genome associated with anorexia nervosa (AN).1 Just like a GPS gives you a range within which your destination point will lie, this study, the Anorexia Nervosa Genetics Initiative (ANGI), has given eating disorder researchers areas on the ‘map’ of the human genome that are likely to hold bits of genetic architecture that contribute to the development of AN. This is exciting because it means researchers now have a well-founded roadmap for further research.
But what will that future research look like?
As mentioned above, this study has provided us with regions but not the specific “co-ordinates,”’ or causal DNA variations. Finding these causal variants is part of the next step in AN genetics research. The roadmap is to focus in on these regions to find the causal variants which will allow us to better understand the biology of the disorder, identify risk markers for early detection and ultimately develop biologically targeted and personalised therapeutics, improving outcome and quality of life for individuals affected by AN.
AN is a highly complex disorder, which means that we anticipate that there will be hundreds if not thousands of genes involved. This is one reason that we need such large numbers of individuals, so that we can even identify genes that have a fairly small effect. In brief this is how the biology of gene expression happens. DNA code is turned into proteins in two stages called ’transcription and translation”. Picture it like the process of taking some notes and writing, editing, and binding it into a book. The gene code (the notes) is transcribed (written out) by a protein called a polymerase, which makes a string of nucleotides complementary to the gene code (imagine the teeth on a zipper). These nucleotides (called RNA) tell the cell machinery which protein to make, or the publisher what words to print. Before the RNA is sent out to the cell machinery, it often undergoes a bit of editing or modification, just like the final edit of a book before it’s published. After this editing, the RNA is sent out into the cell where the cellular machinery translates it into a protein. These proteins then have myriad effects on how our bodies look and function from the cellular level all the way up to visible characteristics like our eye color.
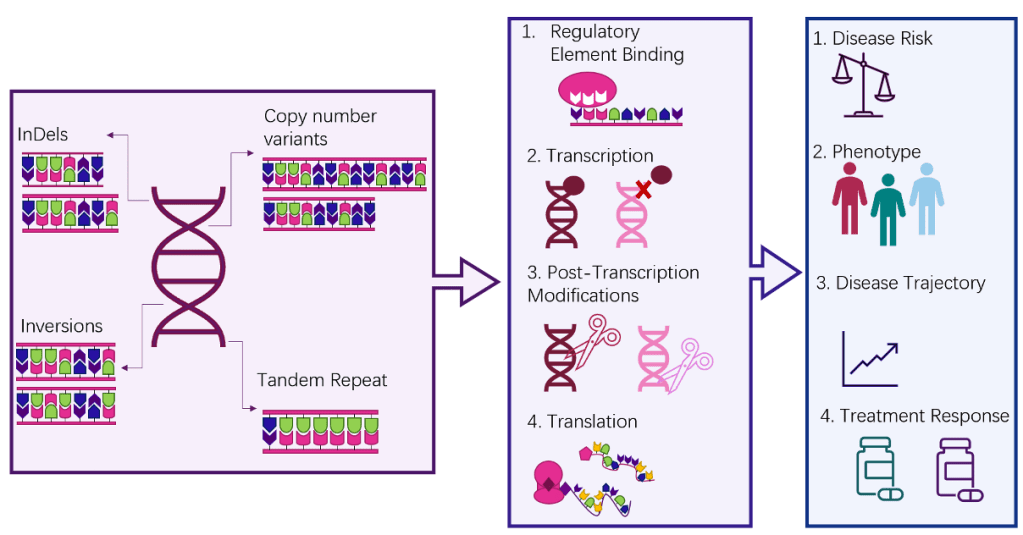
Genetics advances very rapidly. At times it feels like the more we know, the more we know how much we don’t know! We are sure, however, that it is much more complex than “one gene equals one trait”. Traits are affected by how gene expression is regulated, how genes and their products affect one another, the chemical interactions of DNA, variations within a gene code, and how external influences (including the environment) can impact all of these processes. Looking at other complex traits, such as schizophrenia or height, we can get an idea of some of the types of genetic architecture that might be underlying AN. These include single nucleotide polymorphisms (SNPs), large structural variants such as copy number variants and variable number tandem repeats, and short structural variants (SSVs) such as short tandem repeats (STRs). We recently published an article reviewing how the SSV group might contribute to AN.2
This paragraph is for readers who want to go deeper into the biology. SSVs are relatively understudied in comparison to other types of genetic architecture but have recently been gaining traction in other complex traits, like amyotrophic lateral sclerosis (ALS or Lou Gehrig’s disease), schizophrenia, and height.3,4,5 SSVs are a group of genetic variants that vary in sequence type and are between 2-50 base pairs in length. The STR subgroup is of particular interest. STRs are repeated sequence motifs, where motifs can be one base pair up to six base pairs in size. A mononucleotide repeat would have a motif of one base pair, for example a single thymine (T) nucleotide repeated 15 times. A trinucleotide repeat would have a motif of three base pairs, for example cytosine, adenosine, and guanine (CAG), where the motif is repeated six times, making the variant a total of 18 base pairs in length. What makes these variants so interesting is the variety of different effects they can have on gene expression and regulation. STRs have been shown to modulate how genes are stored, the rate and amount of protein they produce, editing of RNA, how the RNA acts out in the cell, and how the RNA interacts with the ribosomes during translation.4
We believe that STR structure and function make them a good candidate for explaining some nuances of the genetic contribution to AN. Part of our current research is identifying and characterising STRs within the eight genomic regions identified in the 2019 ANGI study. In other words, identifying the specific co-ordinates of our destination within the GPS range. Once we have these specific co-ordinates, we will explore how to use them as markers to identify individuals at risk, understand clinical diversity, and course of illness, to unpack the biological pathways underpinning the disorder, and, we hope, to identify specific therapeutic targets based on enhanced understanding of the biology of AN. Although genetic research on other eating disorders is lagging behind, ongoing work in bulimia nervosa, binge-eating disorder, atypical anorexia nervosa, and avoidant/restrictive food intake disorder will position us well to apply these methods to all eating disorders.
References
1. Watson, H. J., Yilmaz, Z., Thornton, L. M., Hübel, C., Coleman, J. R. I., Gaspar, H. A., Bryois, J., Hinney, A., Leppä, V. M., Mattheisen, M., Medland, S. E., Ripke, S., Yao, S., Giusti-Rodríguez, P., Hanscombe, K. B., Purves, K. L., Adan, R. A. H., Alfredsson, L., Ando, T., . . . Bulik, C. M. (2019). Genome-wide association study identifies eight risk loci and implicates metabo-psychiatric origins for anorexia nervosa. Nature Genetics, 51(8), 1207-1214. https://doi.org/10.1038/s41588-019-0439-2
2. Berthold, N., Pytte, J., Bulik, C. M., Tschochner, M., Medland, S. E., & Akkari, P. A. (2022). Bridging the gap: Short structural variants in the genetics of anorexia nervosa. International Journal of Eating Disorders, 55(6), 747-753. https://doi.org/https://doi.org/10.1002/eat.23716
3. Fotsing, S. F., Margoliash, J., Wang, C., Saini, S., Yanicky, R., Shleizer-Burko, S., Goren, A., & Gymrek, M. (2019). The impact of short tandem repeat variation on gene expression. Nature Genetics, 51(11), 1652-1659. https://doi.org/10.1038/s41588-019-0521-9
4. Gymrek, M., Willems, T., Guilmatre, A., Zeng, H., Markus, B., Georgiev, S., Daly, M. J., Price, A. L., Pritchard, J. K., Sharp, A. J., & Erlich, Y. (2016). Abundant contribution of short tandem repeats to gene expression variation in humans. Nature Genetics, 48(1), 22-29. https://doi.org/10.1038/ng.3461
5. Theunissen, F., Anderton, R. S., Mastaglia, F. L., Flynn, L. L., Winter, S. J., James, I., Bedlack, R., Hodgetts, S., Fletcher, S., Wilton, S. D., Laing, N. G., MacShane, M., Needham, M., Saunders, A., Mackay-Sim, A., Melamed, Z. e., Ravits, J., Cleveland, D. W., & Akkari, P. A. (2021). Novel STMN2 Variant Linked to Amyotrophic Lateral Sclerosis Risk and Clinical Phenotype. Frontiers in Aging Neuroscience, 13, 127. https://www.frontiersin.org/article/10.3389/fnagi.2021.658226